pacman::p_load(seriation, dendextend, heatmaply, tidyverse)Hands-on Exercise 5c: Heatmap for Visualising and Analysing Multivariate Data
1 Overview
Heatmaps visualise data through variations in colouring. When applied to a tabular format, heatmaps are useful for cross-examining multivariate data, by placing variables in the columns and observations (or records) in rows and colouring the cells within the table.
Heatmaps are good for showing variance across multiple variables, revealing any patterns, displaying whether any variables are similar to each other, and detecting if any correlations exist in-between them.
This exercise aims to plot static and interactive heatmaps for visualising and analysing multivariate data.
2 Getting Started
2.1 Installing and loading the packages
For this exercise, the following R packages will be used.
Seriation: The package provides several visualizations (grid and ggplot2) to reveal structural information, including permuted image plots, reordered heatmaps, Bertin plots, clustering visualizations like dissimilarity plots, and visual assessment of cluster tendency plots (VAT and iVAT).
dendextend: provides general functions for handling tree-like structures in R.
2.2 Data import
The data of World Happines 2018 report will be used. The data set is downloaded from here.
wh <- read_csv("data/WHData-2018.csv")2.3 Preparing the Data
Let us change the rows by country name instead of row number. And then convert this to a data matrix to make our heatmap.
row.names(wh) <- wh$Country
wh1 <- dplyr::select(wh, c(3, 7:12))
wh_matrix <- data.matrix(wh)3 Static Heatmap
There are many R packages and functions can be used to drawing static heatmaps, they are:
heatmap() of R stats package. It draws a simple heatmap.
heatmap.2() of R ggplots package. It draws an enhanced heatmap compared to the R base function.
pheatmap() of pheatmap R package. pheatmap package also known as Pretty Heatmap. The package provides functions to draw pretty heatmaps and provides more control to change the appearance of heatmaps.
ComplexHeatmap package of R/Bioconductor package. The package draws, annotates and arranges complex heatmaps (very useful for genomic data analysis). The full reference guide of the package is available here.
superheat package. A system for generating extendable and customizable heatmaps for exploring complex datasets, including big data and data with multiple data types. The full reference guide of the package is available here.
We will plot a heatmap by using heatmap() of Base Stats.
By default, heatmap() plots a cluster heatmap.
h_heatmap <- heatmap(wh_matrix,
Rowv=NA, Colv=NA)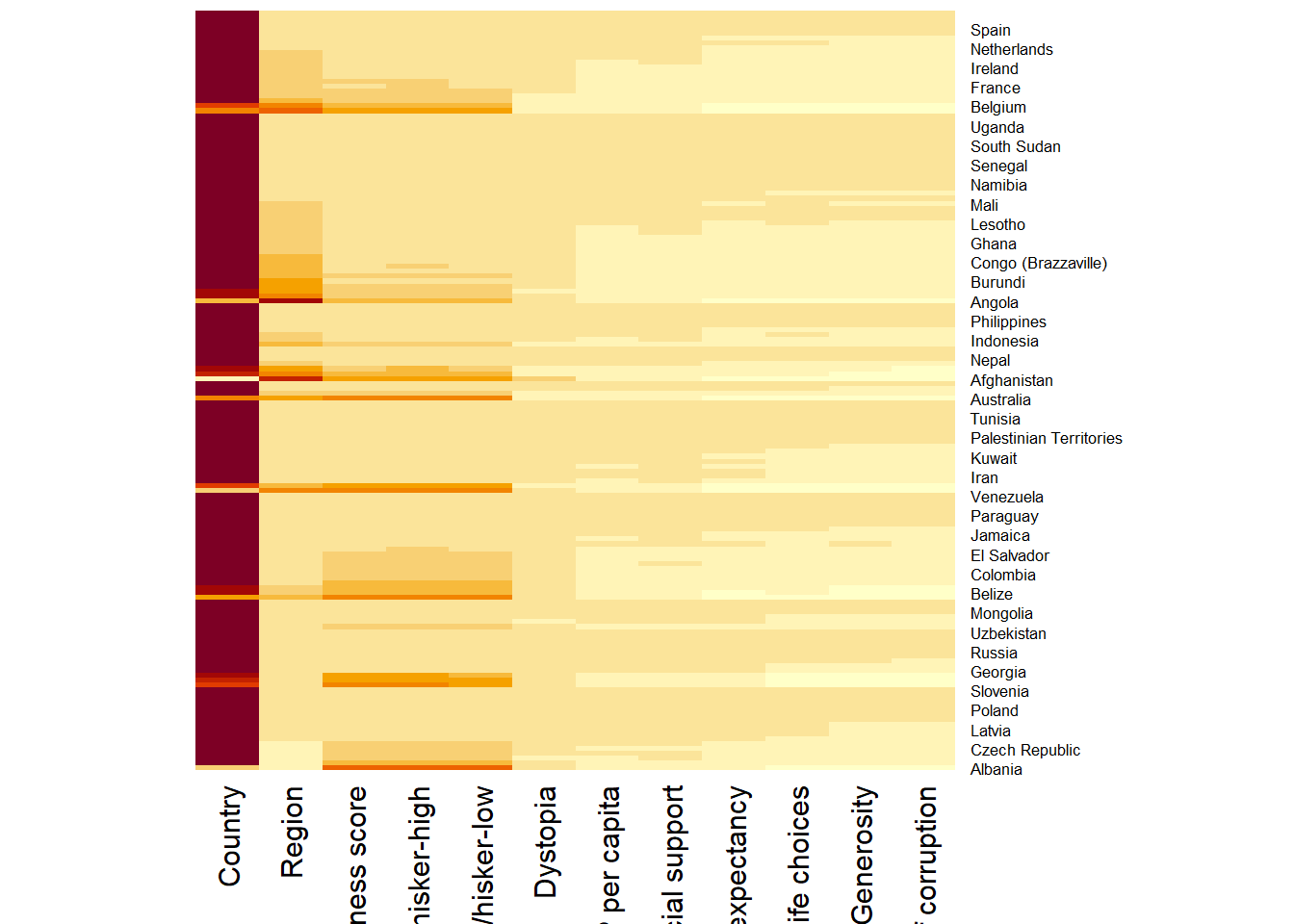
The arguments Rowv=NA and Colv=NA are used to switch off the option of plotting the row and column dendrograms.
wh_heatmap <- heatmap(wh_matrix)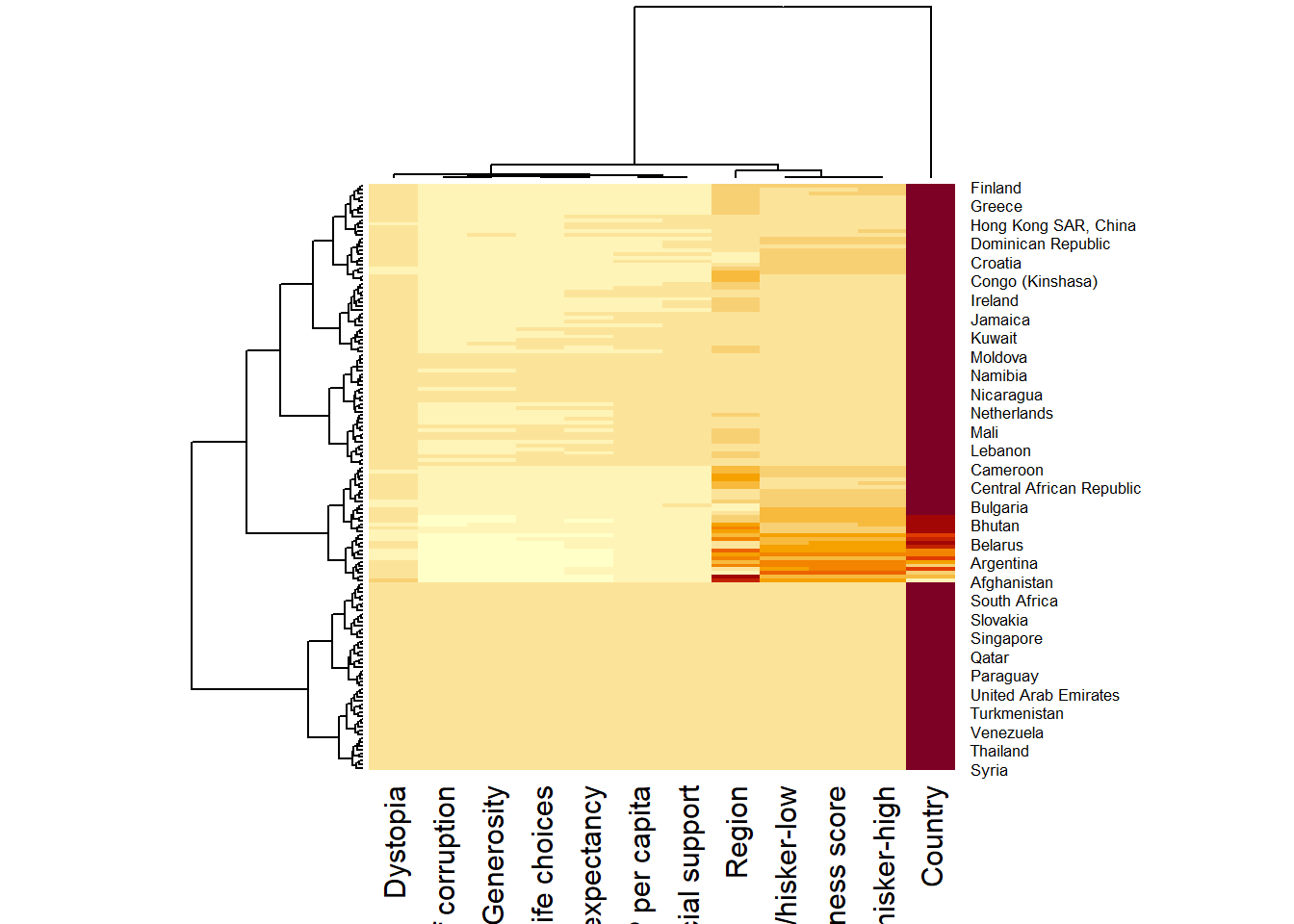
The order of both rows and columns is different compare to the default heatmap. To display the dendrogram, heatmap does a reordering using clusterisation: it calculates the distance between each pair of rows and columns and try to order them by similarity.
Each variable has different ranges and maximum values. Hence, variables with higher values (e.g. Country) will appear in darker colour, which makes other variables’ values insignificant. Therefore, we need to normalize the matrix using the scale argument. It can be applied to rows or to columns.
wh_heatmap <- heatmap(wh_matrix,
scale="column",
cexRow = 0.6,
cexCol = 0.8,
margins = c(10, 4))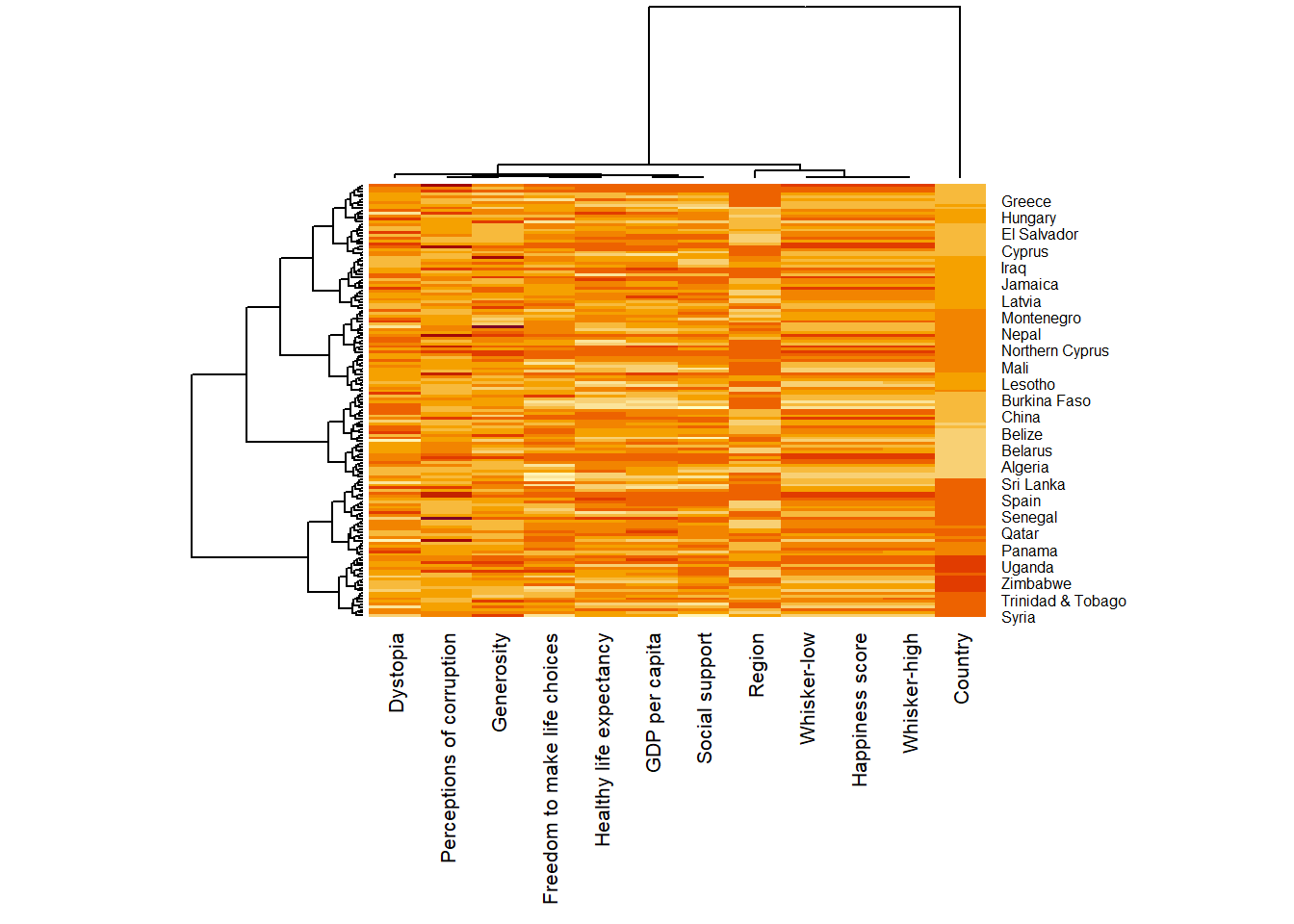
The
marginsargument is used to ensure that the entire x-axis labels are displayed completely.The
cexRowandcexColarguments are used to define the font size used for y-axis and x-axis labels respectively.
4 Interactive Heatmap
heatmaply is an R package for building interactive cluster heatmaps that can be shared online as a stand-alone HTML file. It is designed and maintained by Tal Galili.
Reference on interactive heatmap can be found in Introduction to Heatmaply and user manual.
heatmaply(wh_matrix[, -c(1, 2, 4, 5)])The default horizontal dendrogram is placed on the left hand side of the heatmap.
The text label of each raw is placed on the right hand side of the heat map.
When the x-axis marker labels are too long, they will be rotated by 135 degree from the north.
4.1 Normalising the data
When analysing multivariate data set, it is very common that the variables in the data sets includes values that reflect different types of measurement. In general, these variables’ values have their own range. In order to ensure that all the variables have comparable values, data transformation are commonly used before clustering.
Three main data transformation methods are supported by heatmaply(), namely: scale, normalize and percentize.
When all variables are came from or assumed to come from some normal distribution, then scaling (i.e.: subtract the mean and divide by the standard deviation) would bring them all close to the standard normal distribution.
In such a case, each value would reflect the distance from the mean in units of standard deviation. The scale argument supports column and row scaling.
heatmaply(wh_matrix[, -c(1, 2, 4, 5)],
scale = "column")When variables in the data come from possibly different (and non-normal) distributions, the normalize function can be used to bring data to the 0 to 1 scale by subtracting the minimum and dividing by the maximum of all observations.
This preserves the shape of each variable’s distribution while making them easily comparable on the same “scale”.
heatmaply(normalize(wh_matrix[, -c(1, 2, 4, 5)]))This is similar to ranking the variables, but instead of keeping the rank values, divide them by the maximal rank. This is done by using the ecdf of the variables on their own values, bringing each value to its empirical percentile.
The benefit of the percentize function is that each value has a relatively clear interpretation, it is the percent of observations that got that value or below it.
heatmaply(percentize(wh_matrix[, -c(1, 2, 4, 5)]))4.2 Clustering algorithm
heatmaply supports a variety of hierarchical clustering algorithms. The main arguments provided are:
distfun: to compute the distance (dissimilarity) between both rows and columns. Default is “dist”. The options “pearson”, “spearman” and “kendall” can be used to use correlation-based clustering, which uses as.dist(1 - cor(t(x))) as the distance metric (using the specified correlation method).hclustfun: to compute the hierarchical clustering whenRowvorColvare not dendrograms. Default is”hclust”.dist_method: default is NULL, which results in “euclidean” to be used. It can accept alternative character strings indicating the method to be passed todistfun. By defaultdistfunis “dist” hence this can be one of “euclidean”, “maximum”, “manhattan”, “canberra”, “binary” or “minkowski”.hclust: default is NULL, which results in “complete” method to be used. It can accept alternative character strings indicating the method to be passed tohclustfun. By defaulthclustfunis hclust hence this can be one of “ward.D”, “ward.D2”, “single”, “complete”, “average” (= UPGMA), “mcquitty” (= WPGMA), “median” (= WPGMC) or “centroid” (= UPGMC).
In general, a clustering model can be calibrated either manually or statistically.
This heatmap is plotted by using hierachical clustering algorithm with “Euclidean distance” and “ward.D” method.
heatmaply(normalize(wh_matrix[, -c(1, 2, 4, 5)]),
dist_method = "euclidean",
hclust_method = "ward.D")In order to determine the best clustering method and number of clusters the dend_expend() and find_k() functions of dendextend package will be used.
First, the dend_expend() will be used to determine the recommended clustering method to be used.
wh_d <- dist(normalize(wh_matrix[, -c(1, 2, 4, 5)]), method = "euclidean")
dend_expend(wh_d)[[3]] dist_methods hclust_methods optim
1 unknown ward.D 0.6137851
2 unknown ward.D2 0.6289186
3 unknown single 0.4774362
4 unknown complete 0.6434009
5 unknown average 0.6701688
6 unknown mcquitty 0.5020102
7 unknown median 0.5901833
8 unknown centroid 0.6338734The output table shows that “average” method should be used because it gave the highest optimum value.
Next, find_k() is used to determine the optimal number of clusters.
wh_clust <- hclust(wh_d, method = "average")
num_k <- find_k(wh_clust)
plot(num_k)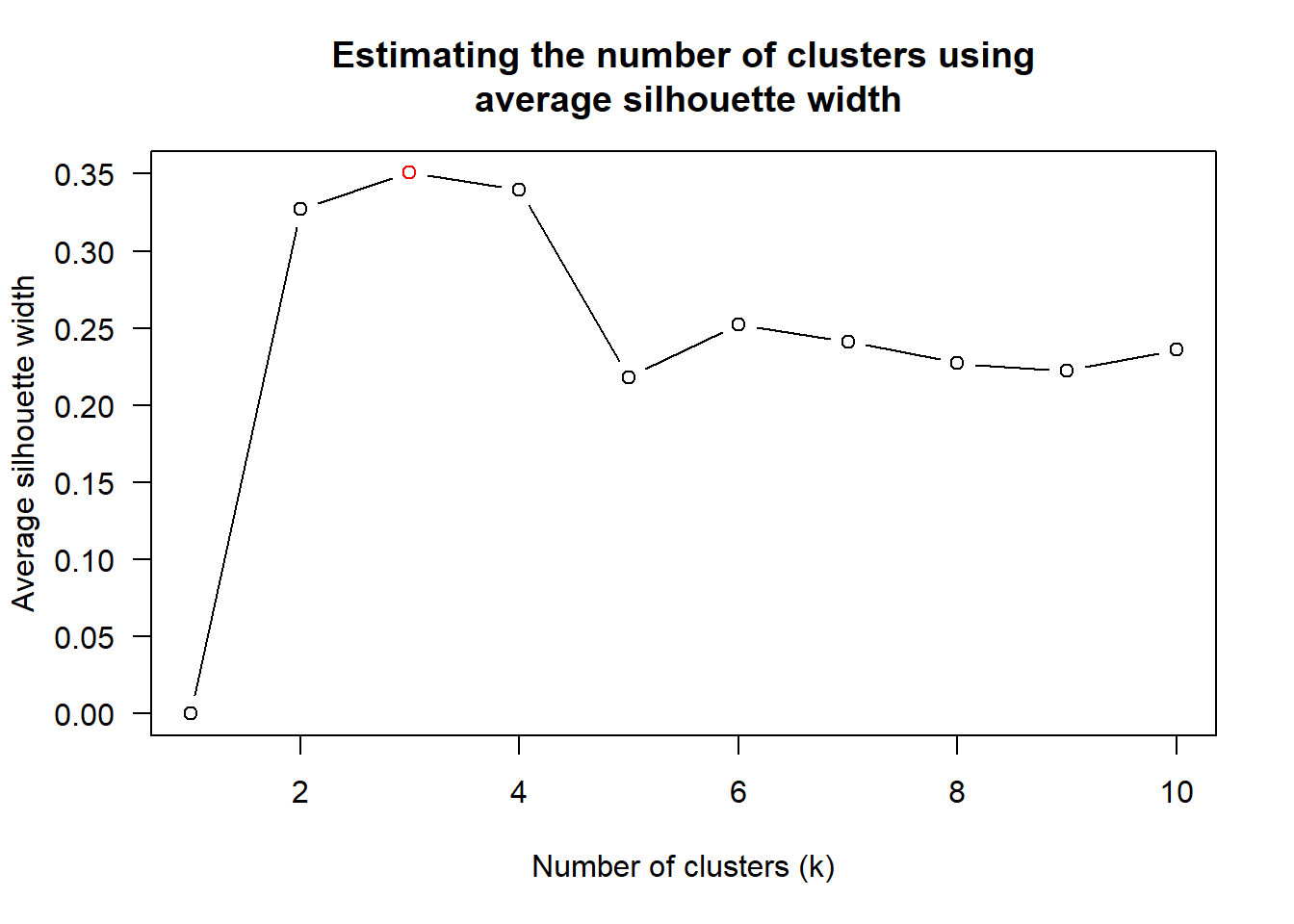
From the graph, the optimal number of clusters, k is 3.
heatmaply(normalize(wh_matrix[, -c(1, 2, 4, 5)]),
dist_method = "euclidean",
hclust_method = "average",
k_row = 3)4.3 Seriation
heatmaply uses the seriation package to find an optimal ordering of rows and columns. Optimal means to optimize the Hamiltonian path length that is restricted by the dendrogram structure. This, in other words, means to rotate the branches so that the sum of distances between each adjacent leaf (label) will be minimized. This is related to a restricted version of the travelling salesman problem.
Optimal Leaf Ordering (OLO), starts with the output of an agglomerative clustering algorithm and produces a unique ordering, one that flips the various branches of the dendrogram around to minimize the sum of dissimilarities between adjacent leaves.
heatmaply(normalize(wh_matrix[, -c(1, 2, 4, 5)]),
seriate = "OLO")“GW” (Gruvaeus and Wainer) which aims for the same goal as OLO but uses a potentially faster heuristic.
heatmaply(normalize(wh_matrix[, -c(1, 2, 4, 5)]),
seriate = "GW")“mean” gives the output we would get by default from heatmap functions in other packages such as gplots::heatmap.2.
heatmaply(normalize(wh_matrix[, -c(1, 2, 4, 5)]),
seriate = "mean")“none” gives us the dendrograms without any rotation that is based on the data matrix.
heatmaply(normalize(wh_matrix[, -c(1, 2, 4, 5)]),
seriate = "none")4.4 Working with colour palettes
The default colour palette uses by heatmaply is viridis. However, we can use other colour palettes in order to improve the aestheticness and visual friendliness of the heatmap. Example of using Blues colour palette of rColorBrewer.
heatmaply(normalize(wh_matrix[, -c(1, 2, 4, 5)]),
seriate = "none",
colors = Blues)4.5 Finishing touch
Beside providing a wide collection of arguments for meeting the statistical analysis needs, heatmaply also provides many plotting features to ensure cartographic quality heatmap can be produced.
In the code chunk below the following arguments are used:
k_rowis used to produce 5 groups.marginsis used to change the top margin to 60 and row margin to 200.fontsizw_rowandfontsize_colare used to change the font size for row and column labels to 4.mainis used to write the main title of the plot.xlabandylabare used to write the x-axis and y-axis labels respectively.
Show the code
heatmaply(normalize(wh_matrix[, -c(1, 2, 4, 5)]),
Colv=NA,
seriate = "none",
colors = Blues,
k_row = 5,
margins = c(NA,200,60,NA),
fontsize_row = 4,
fontsize_col = 5,
main="World Happiness Score and Variables by Country, 2018 \nDataTransformation using Normalise Method",
xlab = "World Happiness Indicators",
ylab = "World Countries")